When people hear the word protein, they often think of chicken or tofu or a protein shake to drink after a workout. Proteins, along with sugars, fats, and carbs, are indeed part of nutrition, but they also carry out many of the functions of life. We eat protein as part of our daily diet, because we cannibalize dietary protein to make our bodies own proteins.
There are many types of proteins—from the enzymes that digest the food in our stomachs, to the structural proteins that allow for cell movement and intracellular transport, to the variety of signals and receptors that guide a cell’s actions and responses to the environment. In the second issue of Hatch, we highlight 25 unique proteins that San Diego scientists are studying to understand biology and treat disease. Below, we give you a bit more about these VIPs.
Becoming a VIP
It is estimated that the human genetic code can produce a million or more different types of proteins. We have dubbed these proteins as “very important proteins” because they are associated with a disease, are a target for a therapy, or a part of a major biological process. Many diseases are caused when proteins are turned on when they should be off. Most cancers, for example, are caused by the inappropriate activation, most often through genetic mutation, of proteins that regulate cell proliferation.
For example, nuclear receptor proteins, SF1 and Nur77, sit in the nucleus, and in response to hormones, they can turn on the genes that support cell proliferation. These proteins are associated with highly aggressive cancers. Biotechs, like Orphagen Pharmaceuticals, work to find either a small molecule or a biologic drug that can target the protein to develop a new cancer therapy.
Proteins can also be made into drugs or therapies. When they are, they fall under the general category of “biologics.” Targeted cancer therapies can be made from modified proteins, such as antibodies. Each antibody is designed to specifically identify a protein or other molecule and disrupt their activity. The side effects of cancer therapies can be significant because the therapies are either generally targeting cells that proliferate or are targeting proteins that are found in both cancer and normal cells.
Other types of proteins can become therapeutics too. Leading Biosciences and OrPro Therapeutics are both creating modified enzymes that could treat shock and a deadly symptom of cystic fibrosis, respectively.
Getting out of shape
When proteins lose their correct shape, they may wreck havoc on a cell, including aggregating together to trigger cell death. Misfolded proteins are the underlying causes of a number of neurodegenerative diseases, including Alzheimer’s disease, Huntington’s disease, ALS, also known as Lou Gehrig’s disease, and prion diseases. Research groups at The Scripps Research Institute (Tainer and Getzoff labs) and UC San Diego (La Spada and Sigurdson labs) are studying how these proteins misfold and how to stop misfolding and the processes to clean up the aggregated proteins.
A protein’s shape is essential to carrying out its function. Like a snowflake, each protein’s shape is unique and often gives clues to the function. However, to the uninitiated eye, very little can be discerned from the arrows and curly ribbons that are used to describe a protein structure. Those that have studied proteins can pick out themes—enzymes, receptors, structural proteins, motor proteins—that each look different to a trained eye. But what happens when structure does not give clues to function?
Looking in a sea of proteins
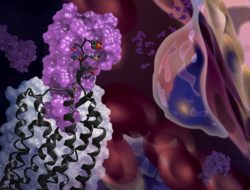
Proteins may be shown as space-filling blobs showing the protein’s approximate shape and interactions with other molecules. This can be especially helpful in designing new drugs that can disrupt these interactions. However, these models present an isolated picture, like a model on a runway. When in fact, the environment inside a cell is packed tighter than a Hong Kong subway car at rush hour.
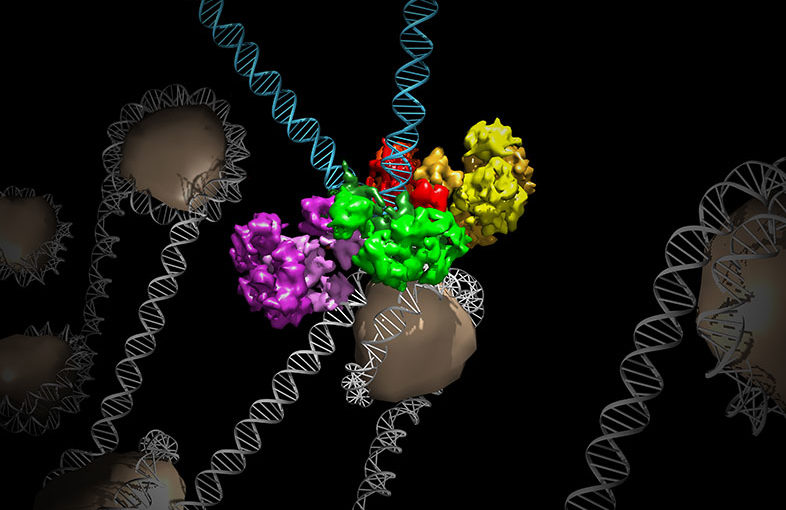
Because the cell has so many proteins and other molecules operating all at once, scientists have to run a number of different tests to interrogate a protein’s function. Sort of like a research game of Clue, the questions revolve around where, with whom, and what happens. Scientists can do yes/no, black-and-white tests that can find a protein’s partners in crime. The tests can say that Professor Lis1 was in the cytoplasm with dynein, but that is circumstantial evidence. A real-time video could give much more information about what actually happened to kill the cell.
Almost every biologist can pick out the shape of GFP (green fluorescent protein), for which the Nobel prize was co-awarded to UC San Diego scientist Roger Tsien, Ph.D. The distinctive barrel-like structure is responsible for the bioluminescence of certain jellyfish. Because the self-contained molecule can light up without any apparent external energy source and be attached to almost any protein, GFP has played a significant role in biological research by real-time tracking of a protein’s location, movement and interactions inside the cell. The GFP technology is also being advanced by local biotech, Avelas Biosciences, to create a real-time “cancer illuminator,” designed to help surgeons distinguish tumor tissue from normal during surgery.
Secrets from a real housekeeper
Scientists can spend most of their careers dissecting what a single protein or a family of proteins do. If the protein is involved in human disease, they may create an animal model with the specific mutation and see if the disease is recapitulated. However, if the protein’s function is not conserved or the same from animal to human, the animal model may provide little information. If it is conserved, scientists will start to characterize the model and test the biology. They can start with understanding protein expression—asking in which tissues and and at what times during life, from embryo to adult, is the protein present and active.
This kind of meticulous analysis unlocked a surprising discovery of what many considered was a boring house-keeping protein, which is a protein that every cell has that does the routine work, like make other proteins or replicate DNA. The lab of Paul Schimmel, Ph.D., and Xiang-Lei Yang, Ph.D., at The Scripps Research Institute found that this particular family of well-conserved, house-keeping proteins were not expressed in a pattern expected of their universal function as part of the protein-making machinery. Indeed, these proteins seemed to have evolved the capability to coordinate the activity of many different cell types including immune cells to restore a stressed or diseased tissue to a healthier state. Today, that discovery is being advanced in the clinic by aTyr Pharma to develop new therapies for rare diseases.
The resistance fighters
Penicillin, the first and probably the most famous antibiotic, was discovered in 1928. Sir Alexander Fleming noticed that mold had contaminated a Petri dish of (Staphylococcus) bacteria, but where the mold was, the bacteria was not. He hypothesized that the mold released a substance that killed the bacteria, and he eventually isolated the substance and called it penicillin. More than a story of how chance favors a prepared mind, this also shows that biology itself can have elegant solutions to human diseases.
Scientists that study infectious diseases and antibiotic resistance are working to attack the problem from multiple sides from blocking the infectious agent from binding to the host’s receptor and entering the cell to targeting a virus’ own protective shell. These include research groups at UC San Diego (Handel lab, Nizet lab), The Scripps Research Institute (Lerner lab, Ollmann Saphire lab, and Burton Ward and Wilson labs) Salk Institute (Lyumkis lab) and La Jolla Institute for Allergy and Immunology (Kronenberg lab).
A fortunate part about developing anti-infectives, including antibiotics, anti-virals and vaccines, is that the proteins these medicines are targeting are foreign. Thus, the side-effect profile of these interventions is often minimal. However, we are in need of new anti-infectives, especially antibiotics, as drug resistance is not an if but a when.
This is where the possibility of making proteins with more than the 20 naturally occurring amino acids could make a big impact. Synthetic biology company, Synthorx, is using an expanded genetic alphabet to create proteins with multiple synthetic amino acids that could be the basis for new antibiotics and better therapeutics.
So now when you look at that big beef or tofu steak, you will know it’s protein, but it’s also going to a be made into a lot of different proteins – and that’s the circle of (protein) life.
A protein by any other name…
The general categories of human proteins and some familiar and not-so familiar representatives.
Enzymatic proteins: protease, lactase, alcohol dehydrogenase, SOD
Defense proteins: antibodies, CD1, thrombin
Receptor proteins: GPCR, CXCR4, integrin
Channel proteins: TRPV2, acetylcholine receptor
Signaling proteins: RhoA, MAPK, FGF1
Hormonal proteins: insulin, prolactin, oxytocin
Storage proteins: ovalbumin, ferritin, casein
Transport proteins: hemoglobin, albumin, cytochromes
Nuclear receptors: Nur77, SF1, androgen receptor
Transcription factors: p53, c-Myc, MECP2
Structural proteins: collagen, keratin, elastin
Movement proteins: dynein, microtubules, TTLL7
Contractile proteins: actin, myosin, dystrophin
Disclosure: Jessica Yingling, Ph.D. is founder and president of Little Dog Communications, whose clients include aTyr Pharma, Avelas Biosciences, Novoron Bioscience, Synthorx.